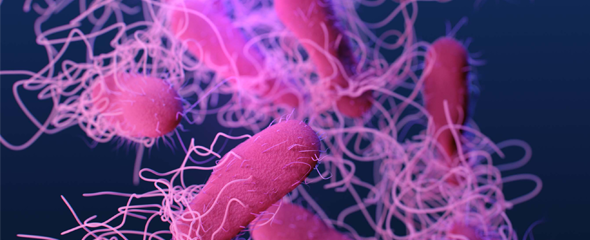
The flagella, which is comprised of tens of thousands of flagellin proteins, is not only important for the movement of Salmonella but also plays a key role in its ability to invade the human host cell. The parts of the flagellin proteins that are exposed are highly versatile and subject to modification by enzymes. One such modification, known as methylation, occurs when a methyl group is added to a protein thus giving the protein the ability to function. Protein methylation plays an integral role in physiological function and signaling pathway modulation in Salmonella enterica.
In the flagella, residues of the amino acid lysine on the flagellin outer surface are methylated to ɛ-N-methyl-lysine. In a previous paper, the researchers led by Prof Michael Kolbe, head of the HZI department “Structural Infection Biology” at CSSB, showed the flagellin’s importance in host cell innovation by demonstrating that flagellin methylation facilitates the adhesion of Salmonella to host cell surfaces. Although it was the first methylase ever discovered more than fifty years ago, little is known about FliB, the methylase responsible for catalyzing this process.
In the current study, Kolbe and his collaborators have revealed that FliB catalyzes a methyl transfer reaction, which is mediated by an iron-sulfur cluster, [4Fe-4S]. According to the researchers, their study sets the stage for understanding the roles of iron-sulfer clusters and radical chemistry in flagellar modification. They were able to demonstrate that FliB is a S-adenosylmethionine dependent methyltransferase. "We also observed that FliB is located in the bacterial cytoplasm,” says the paper’s first author Chu Wang. “This indicates methylation happens prior to flagellin secretion and flagellar assembly.”
“The new insights generated into FliB have helped us characterize a key virulence factor of Salmonella enterica,” explains Michael Kolbe, the paper’s corresponding author. “Although FliB has a low sequence conservation, it is present in many, many different bacterial pathogens and could therefore be an interesting target for future studies.”
Press release on the website of CSSB
Original publication:
Wang C, Nehls C, Baabe D, Burghaus O, Hurwitz R, Gutsmann T, Bröring M, Kolbe M. (2021) Flagellin lysine methyltransferase FliB catalyzes a [4Fe-4S] mediated methyl transfer reaction. PLoS Pathog.17(11):e1010052. doi: 10.1371/journal.ppat.1010052