Veranstaltungen
Am HZI-Campus und den Standorten des HZI finden regelmäßig Veranstaltungen für verschiedene Zielgruppen statt. Hier finden Sie eine regelmäßig aktualisierte Liste mit Informationen über anstehende Events wie Vorträge, wissenschaftliche Symposien, Preisverleihungen und Veranstaltungen für die allgemeine Öffentlichkeit.
HZI Forum
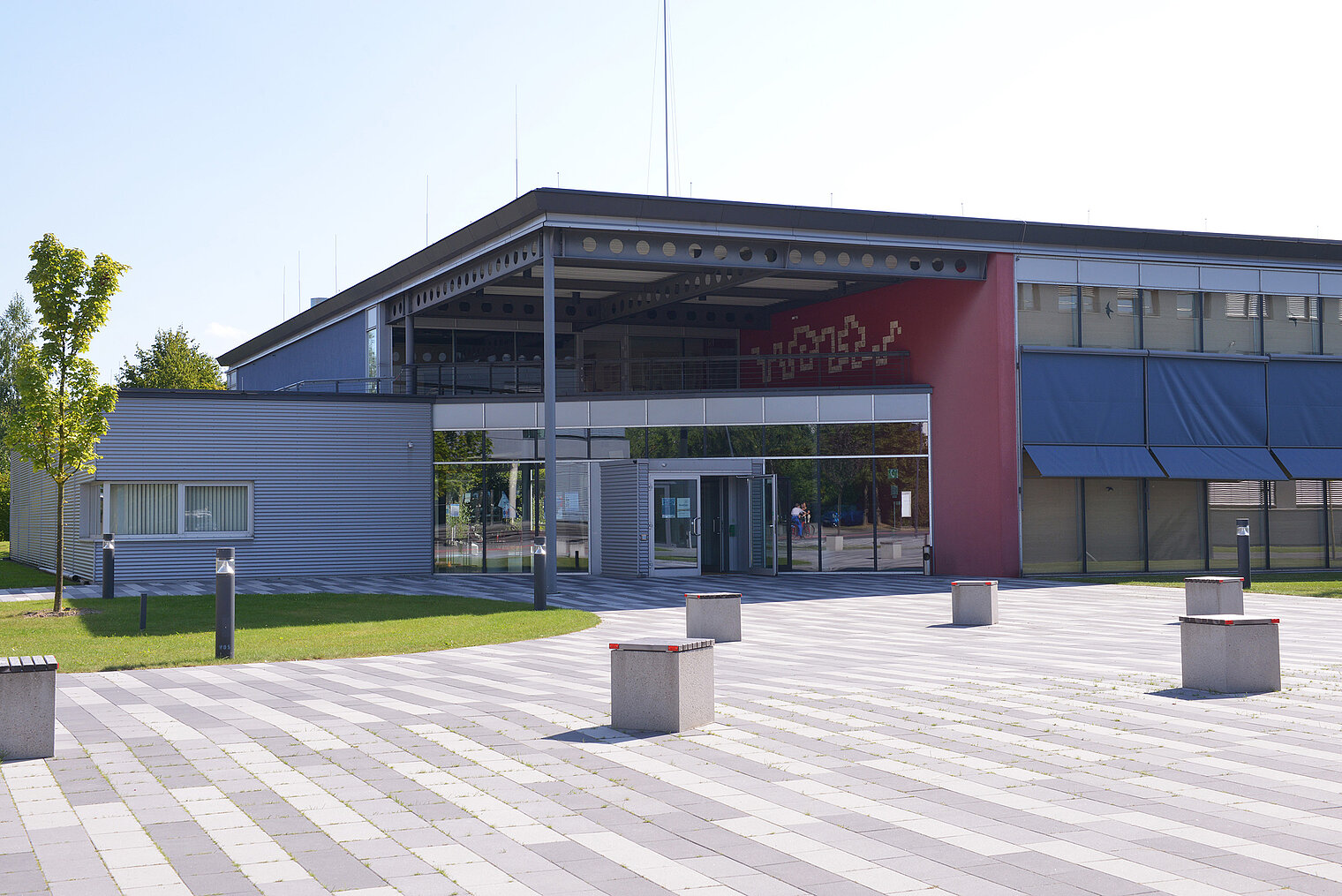

Das Helmholtz-Zentrum für Infektionsforschung hat im Jahr 2000 das HZI Forum als Versammlungsstätte für Vorträge, Seminare, Events und Symposien auf dem Campus eröffnet. Die Räume können auch von externen Unternehmen, Verbänden und anderen Interessenten mit wissenschaftlichem Hintergrund angemietet werden.
Zur Verfügung stehen ein Saal mit Bühne (X0.13), zwei große Seminarräume (X0.13a, X1.04) und ein kleiner Seminarraum (X1.01) sowie dazu ein Foyer, eine Galerie und eine begrünte Außenfläche. Hier finden Sie Raumpläne mit Grundbestuhlung und Alternativbestuhlung.
Sollten Sie sich für eine Anmietung eines oder mehrerer Räume interessieren, freuen wir uns über Ihre Kontaktaufnahme.

Tag der offenen Tür am HZI
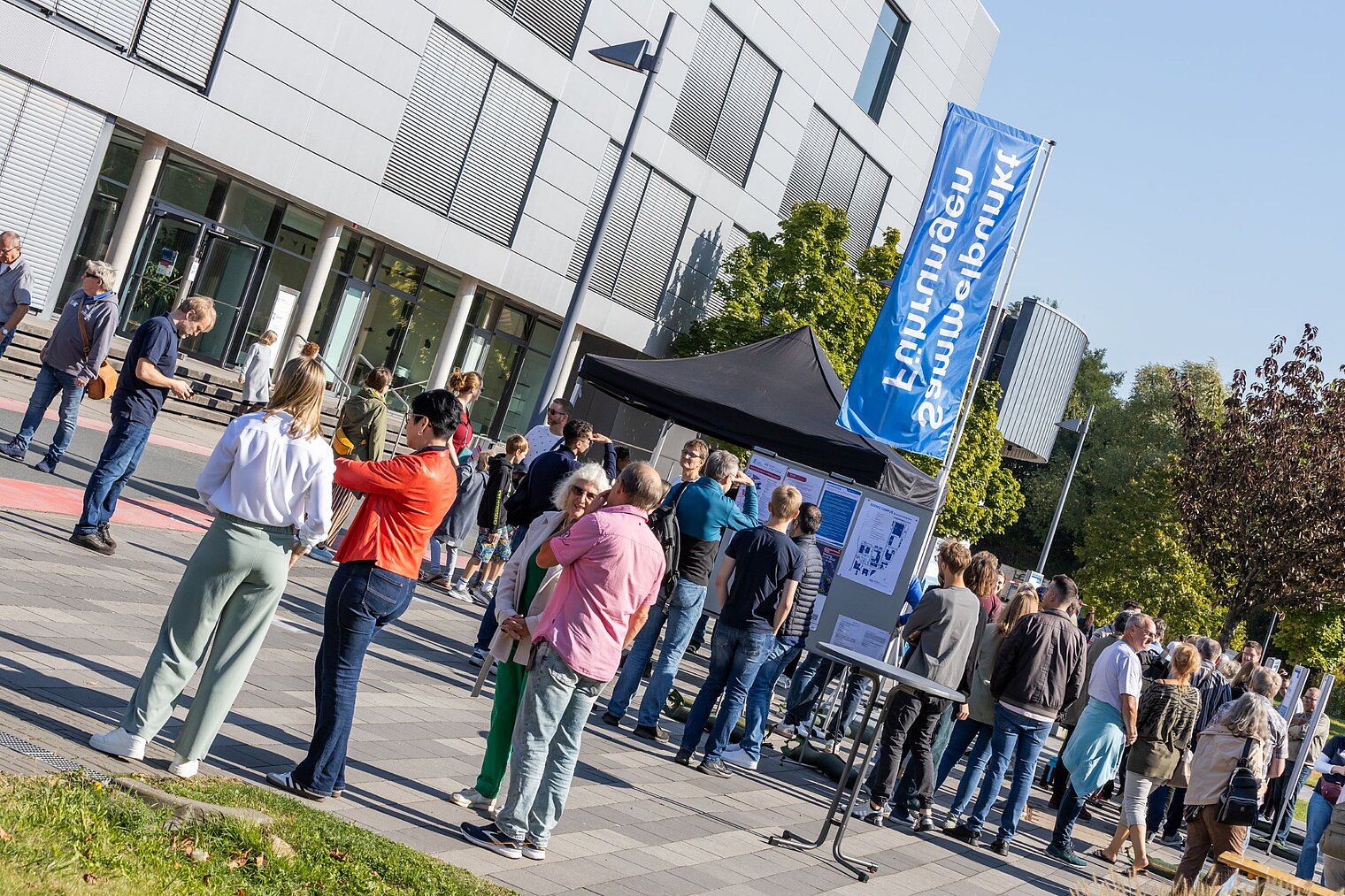
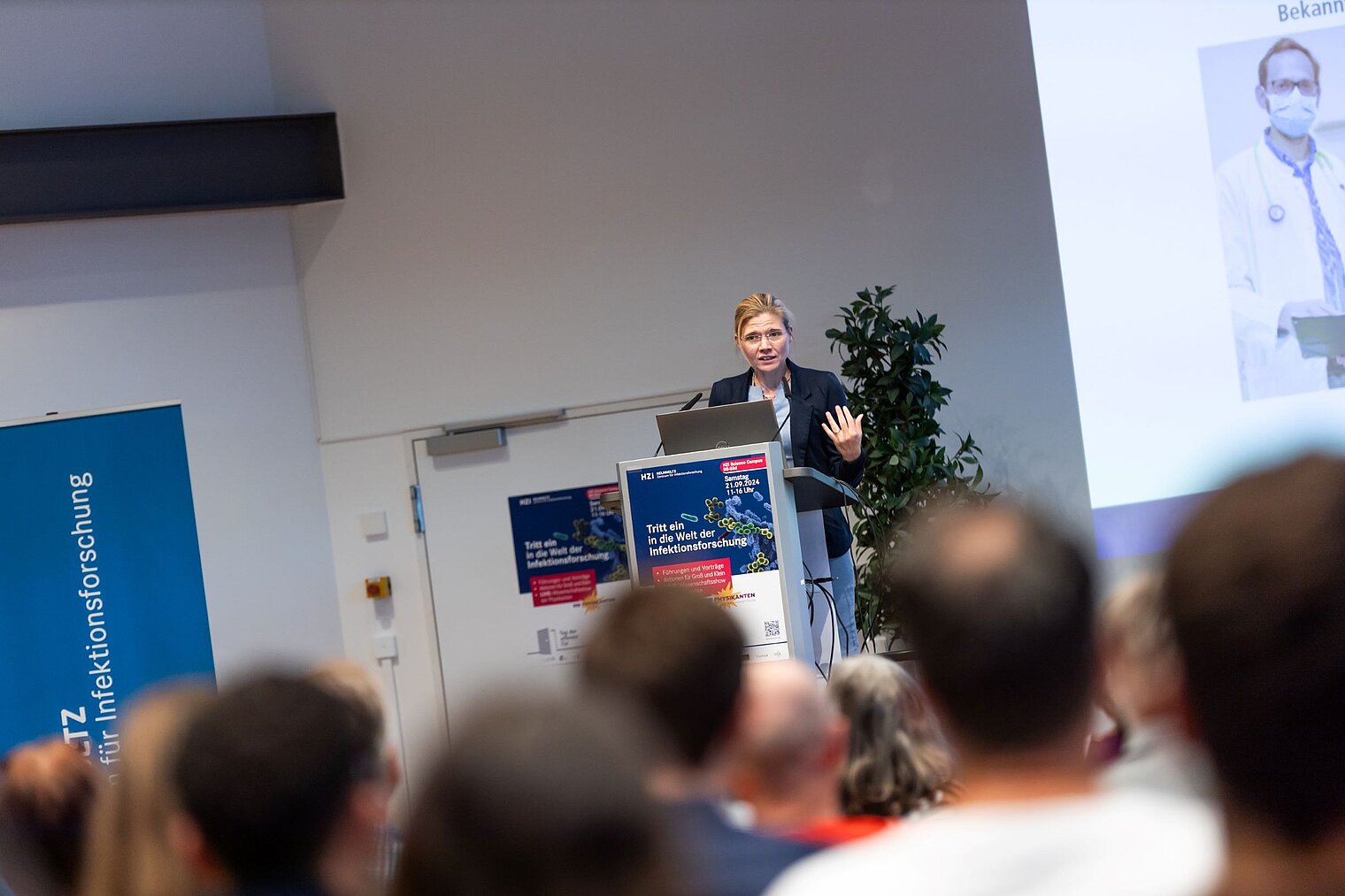
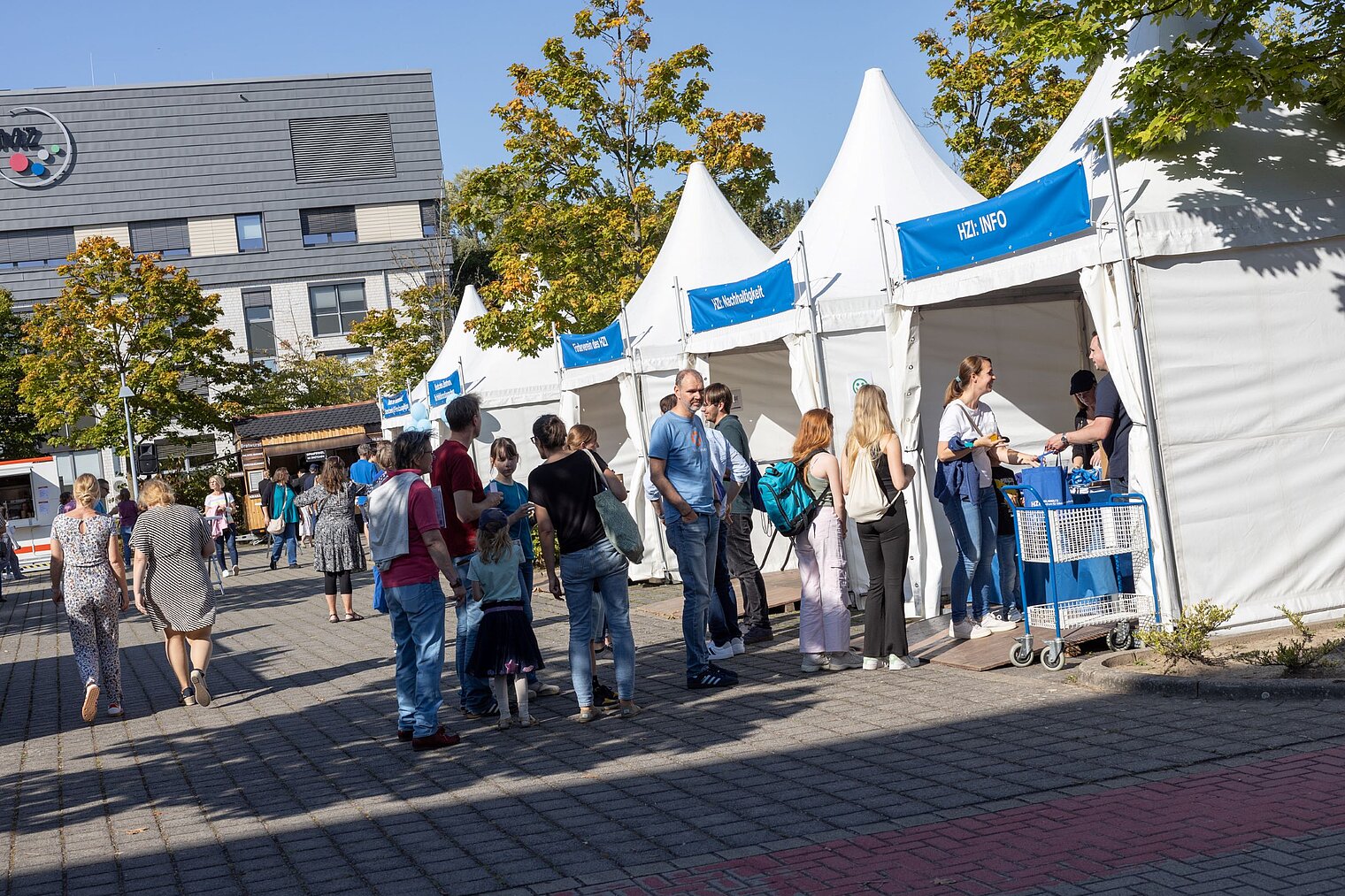
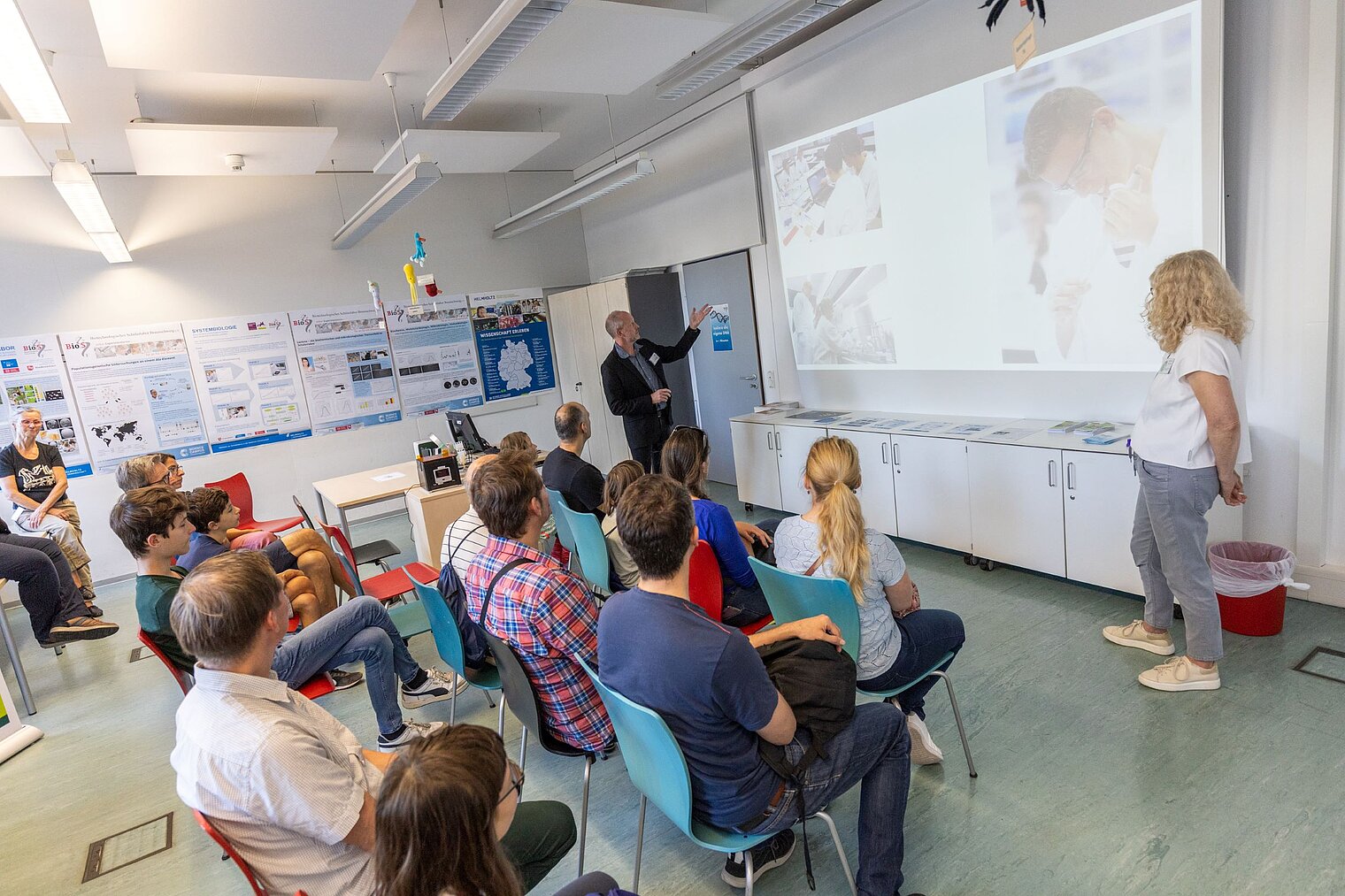
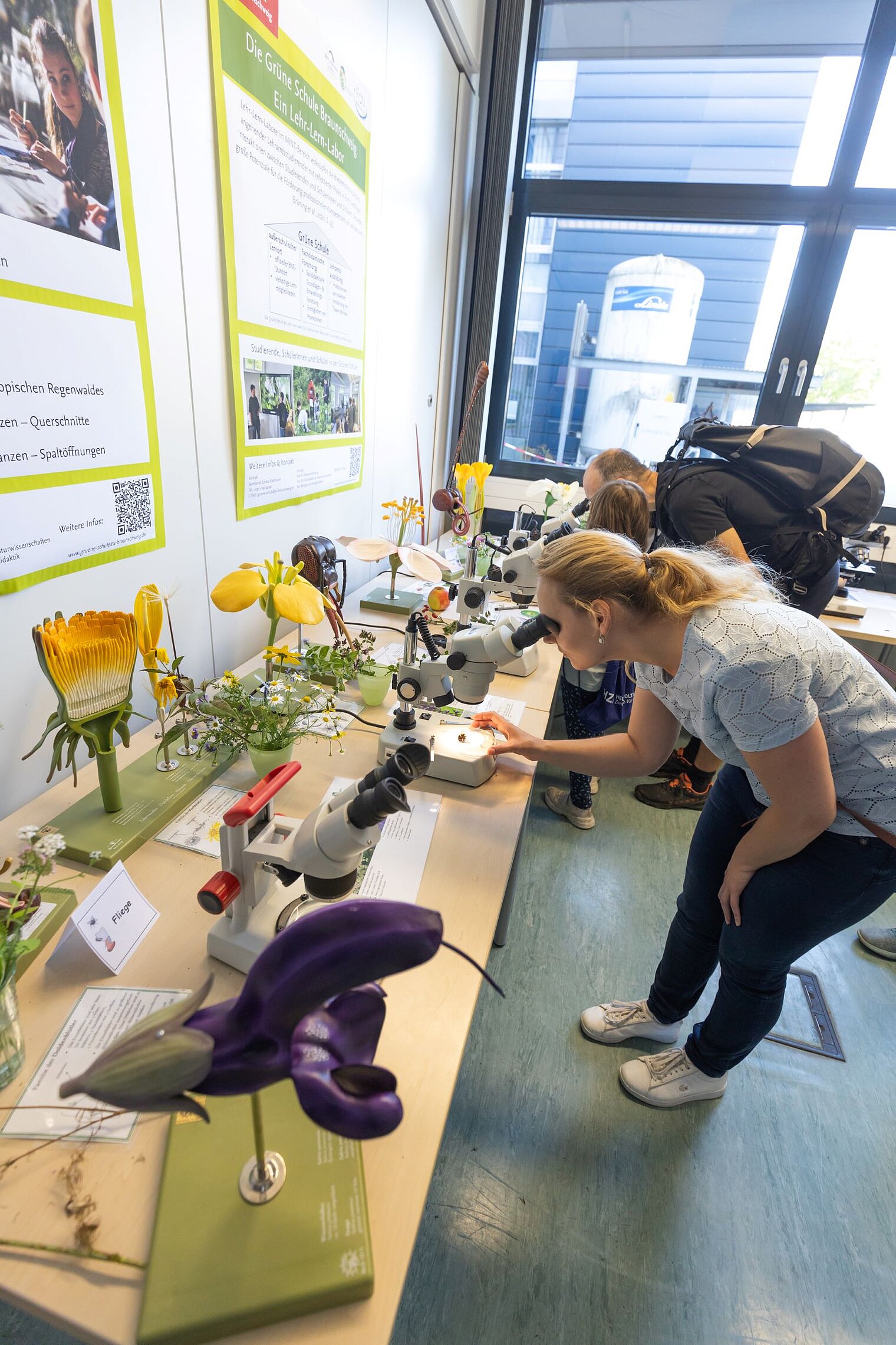
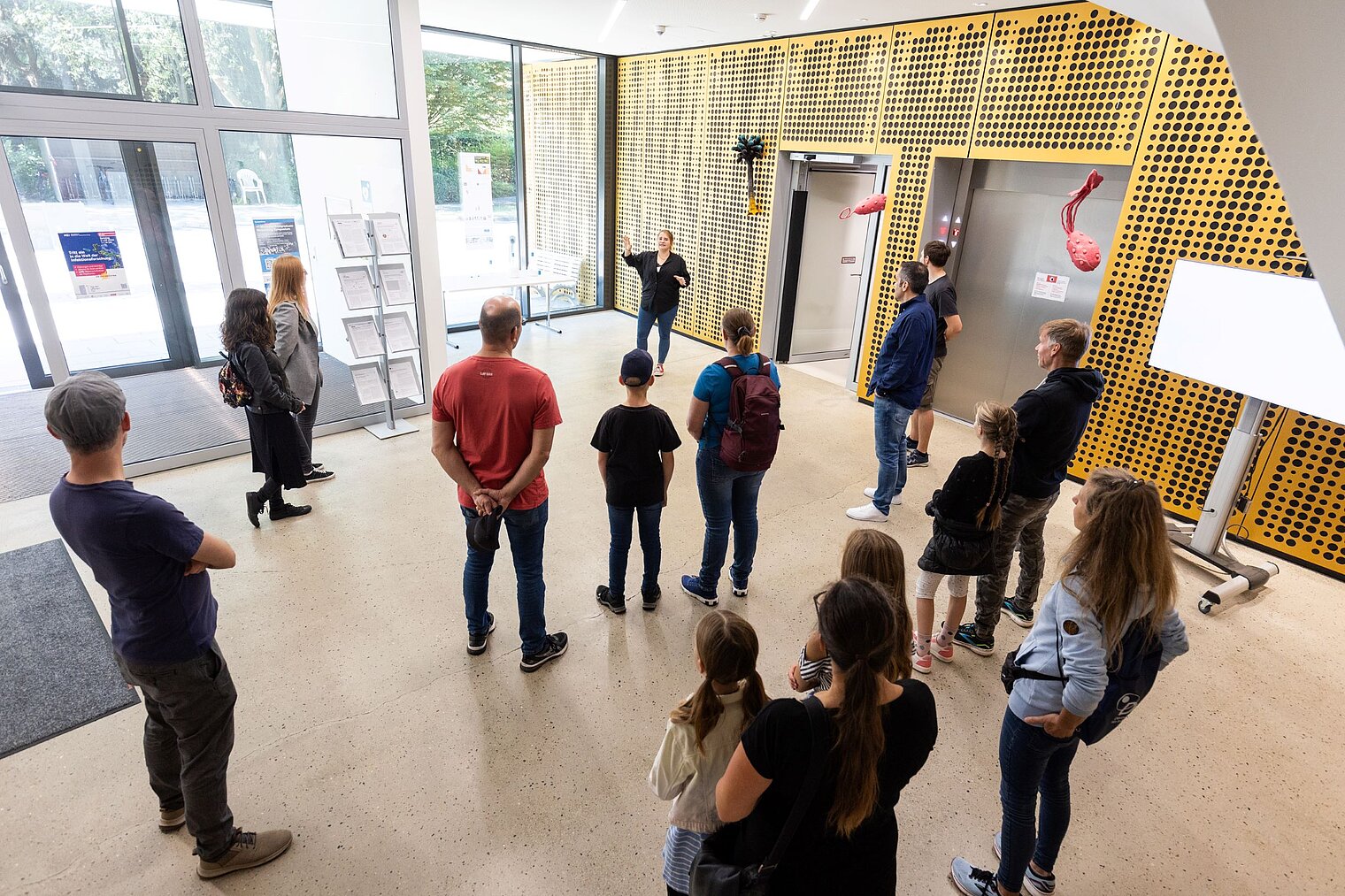
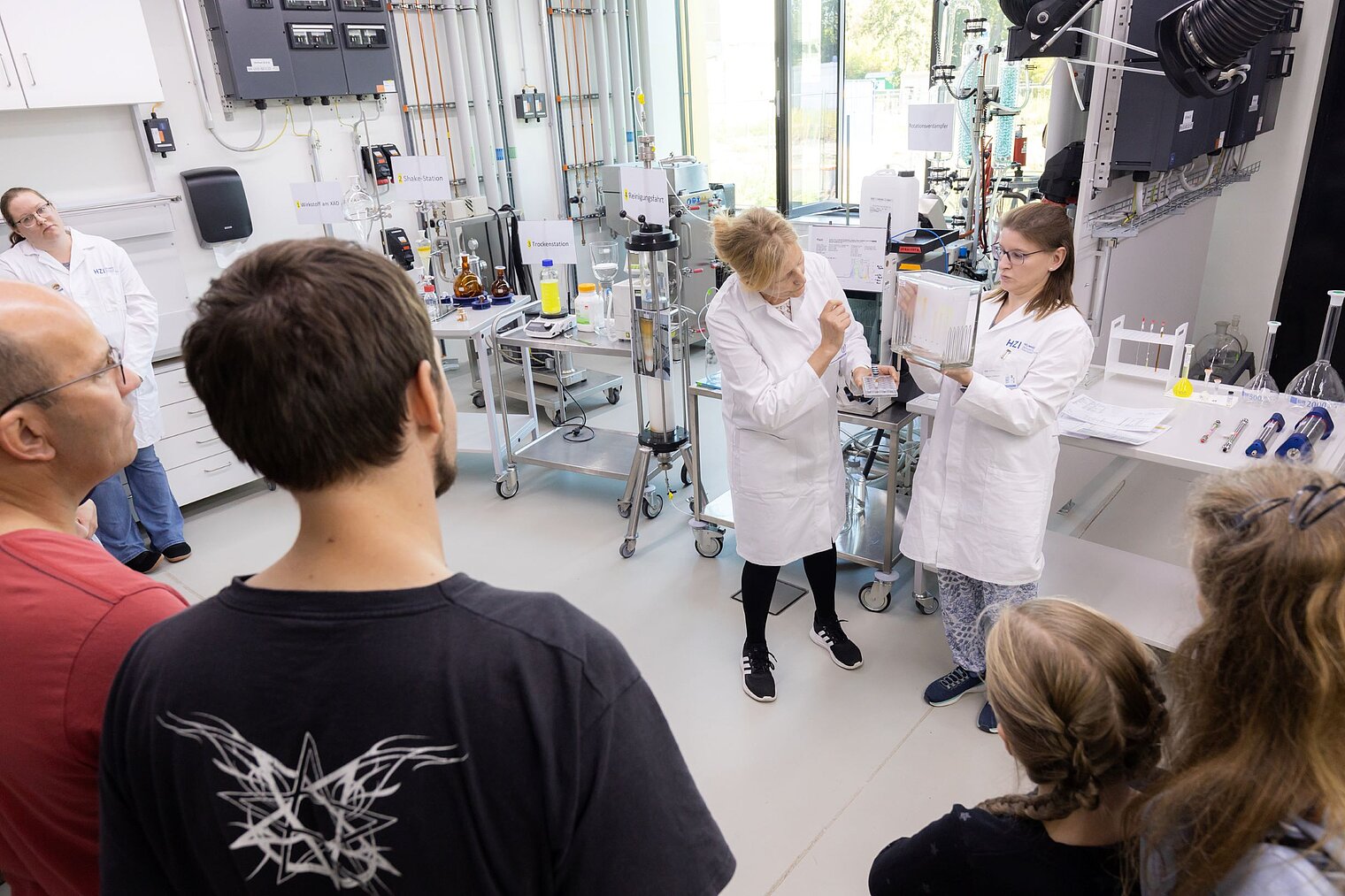
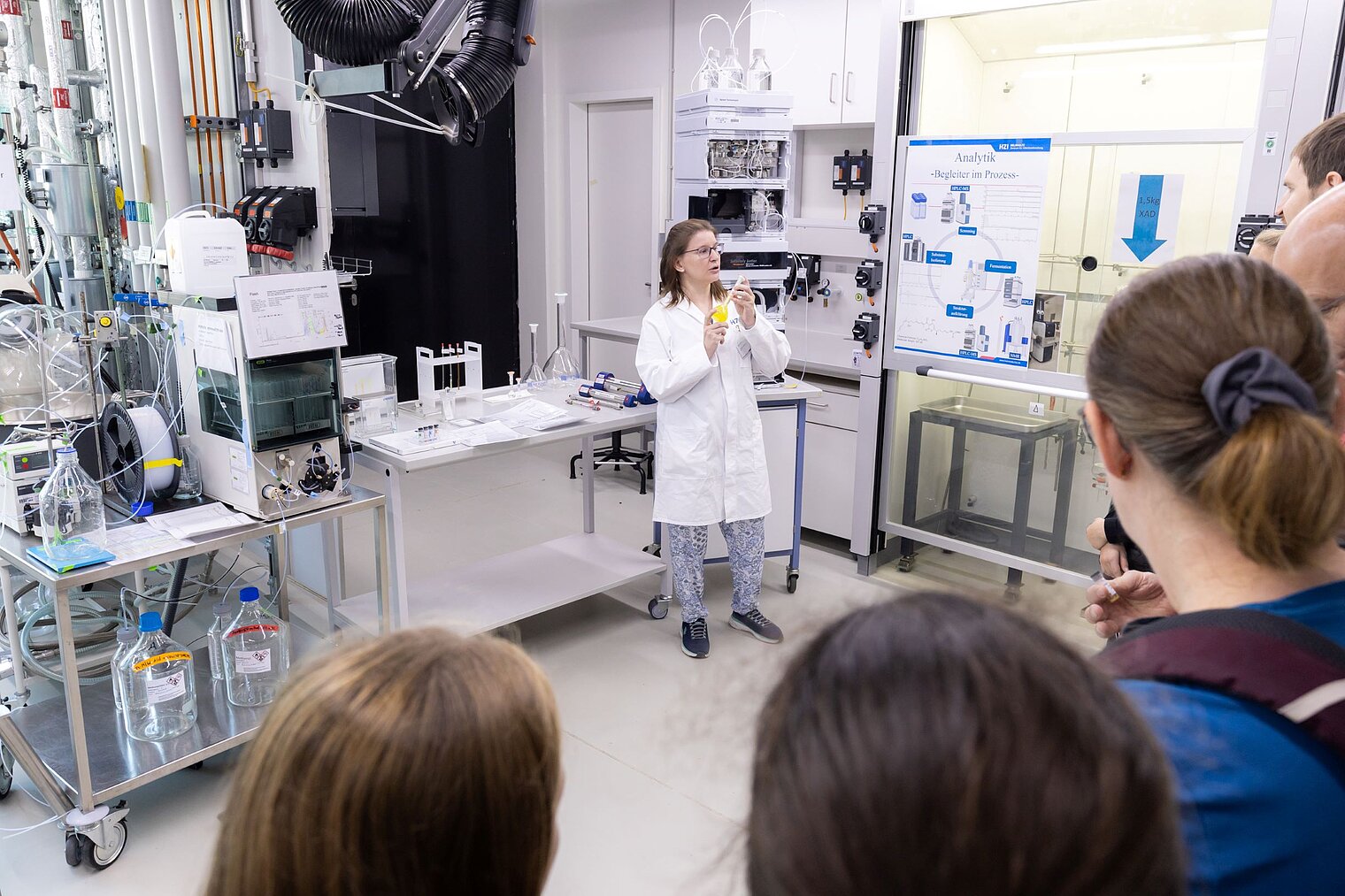
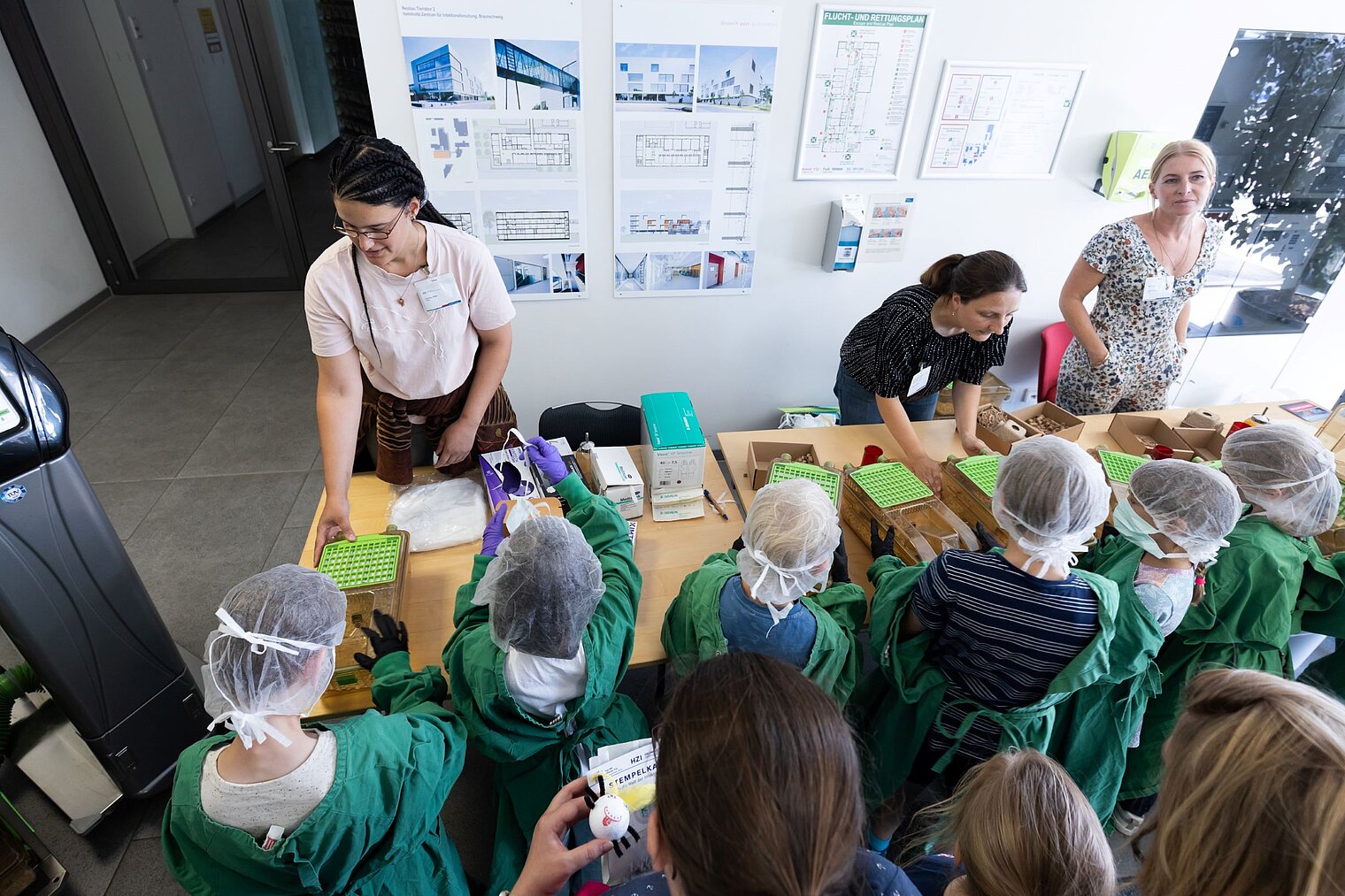
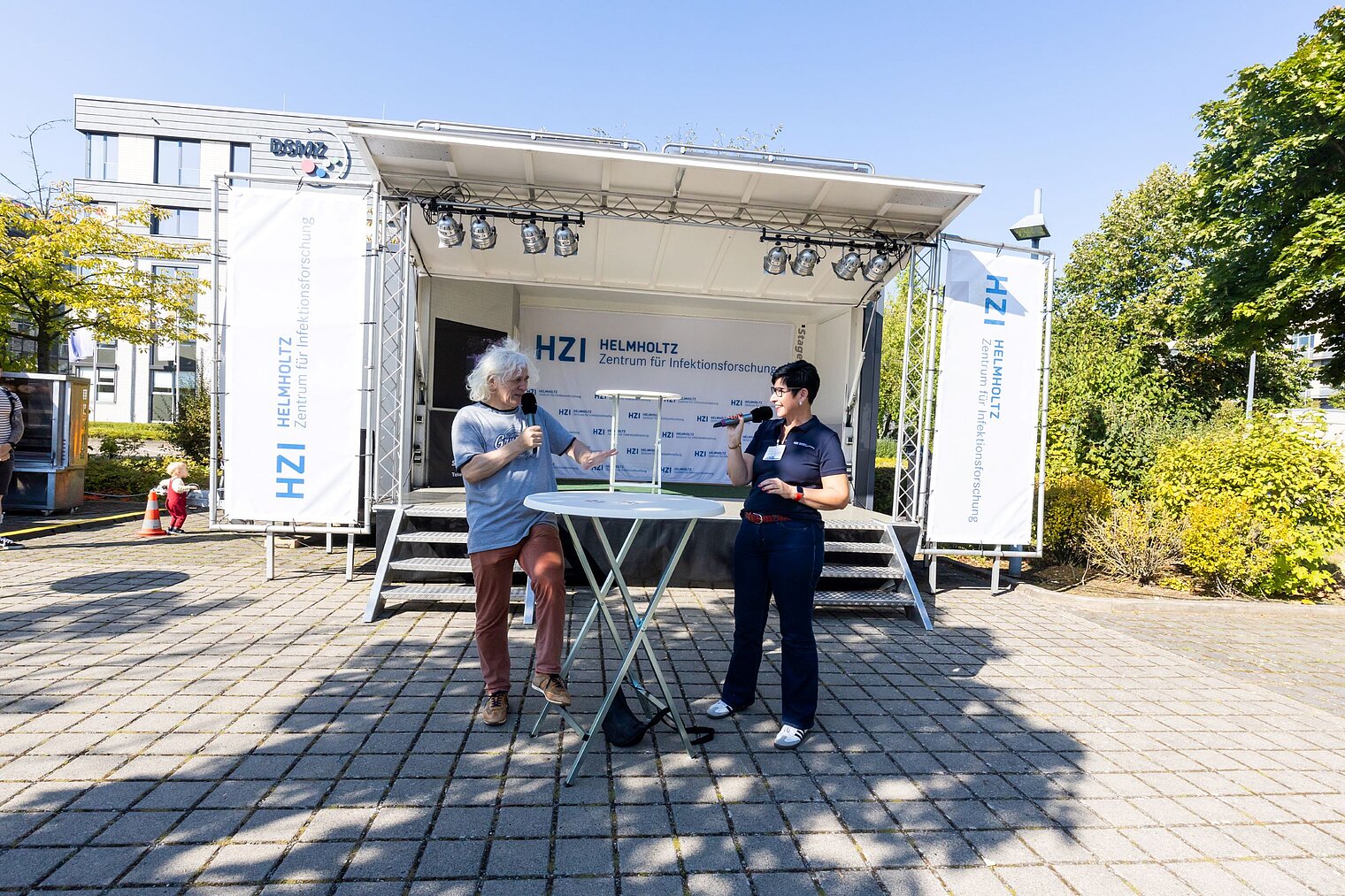
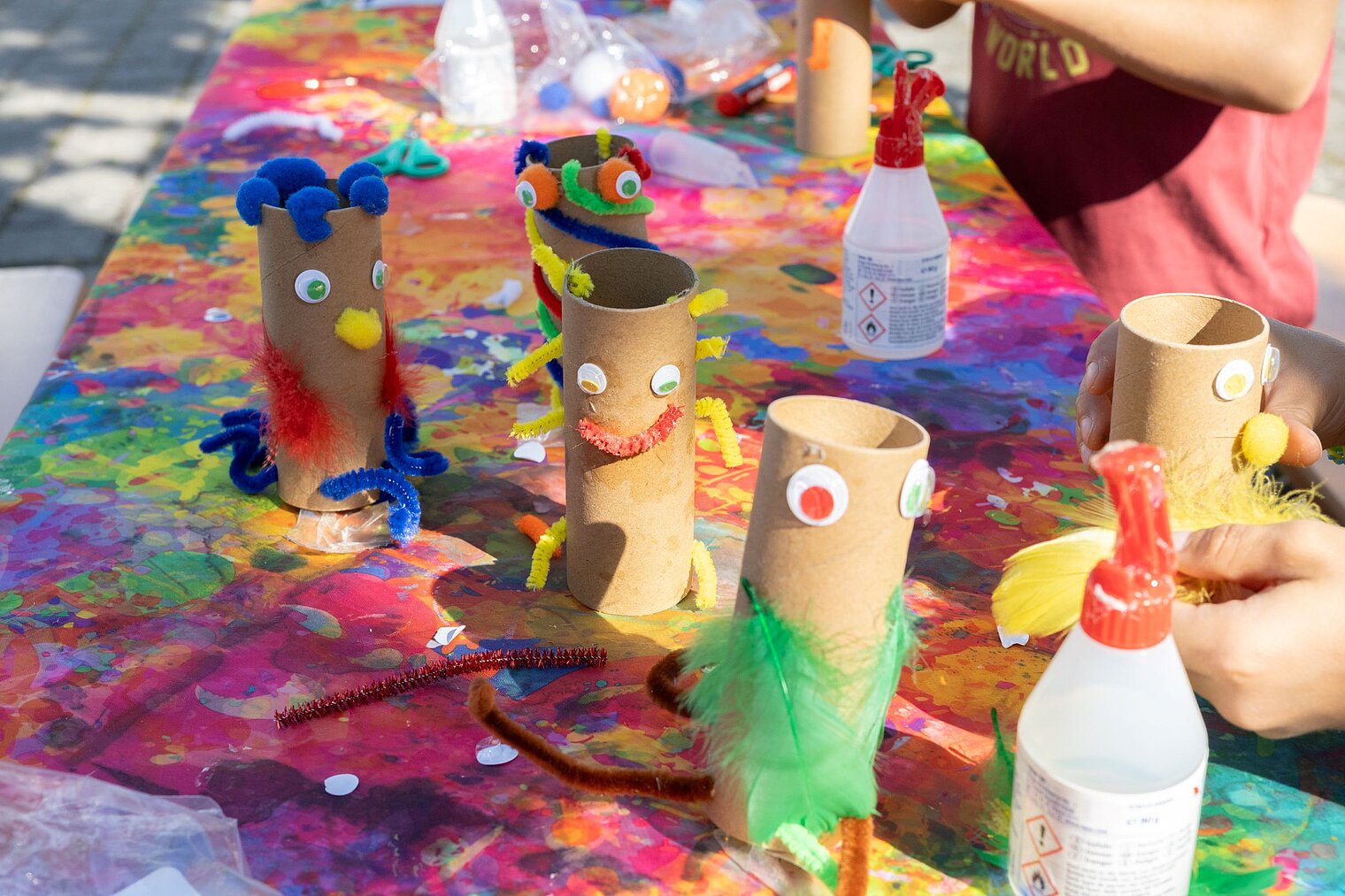
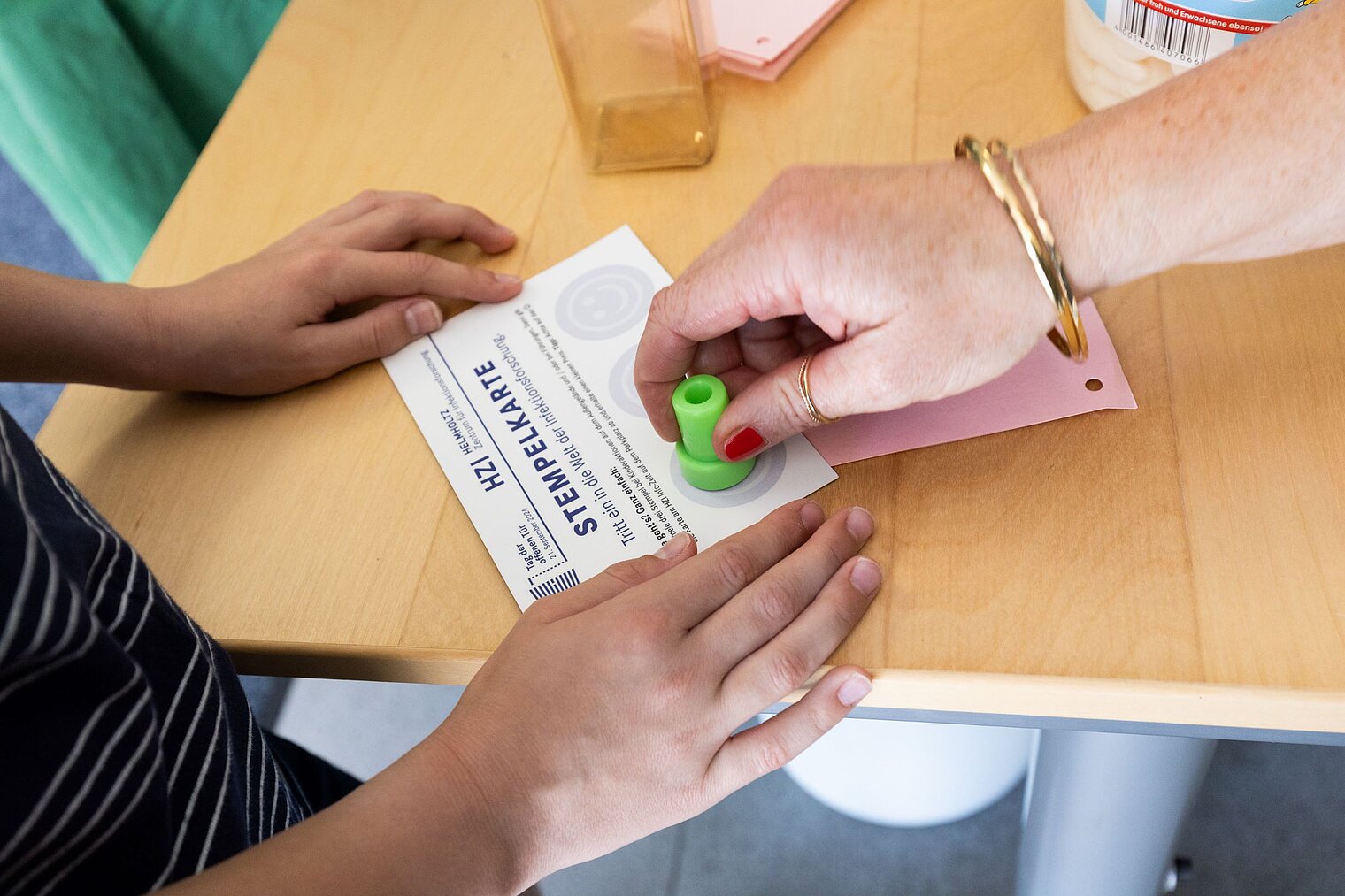
Echte Labore besichtigen, mit Wissenschaftler:innen über ihre Arbeit sprechen und dazwischen noch Vorträge von Expert:innen zu spannenden Themen aus ihrer Forschung hören: Der Tag der offenen Tür am HZI ist eine einzigartige Möglichkeit, in die Welt der Infektionsforschung einzutreten! Dadurch wird das volle Spektrum unserer Forschungsthemen erlebbar. Zusätzlich bieten wir ein buntes Rahmenprogramm für alle Altersgruppen an. Auch die kleinen Besucher:innen kommen durch das abwechslungsreiche Kinderprogramm voll auf ihre Kosten.
Der Tag der offenen Tür des HZI im September 2024 wurde von ca. 1300 Personen besucht. Über zukünftige Termine (voraussichtlich im Jahr 2026) werden wir rechtzeitig auf der HZI-Webseite informieren.
Musik und Infektionen
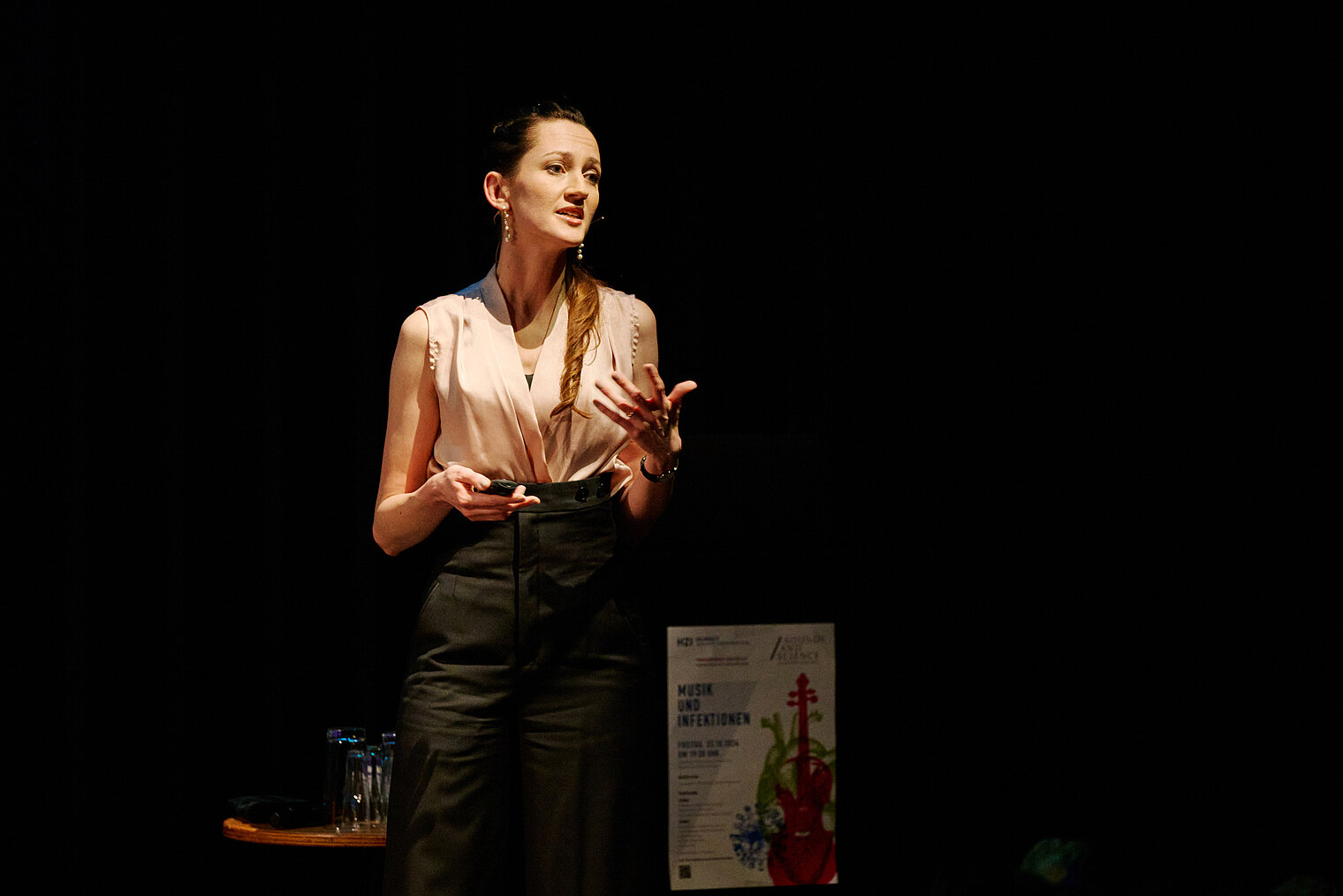
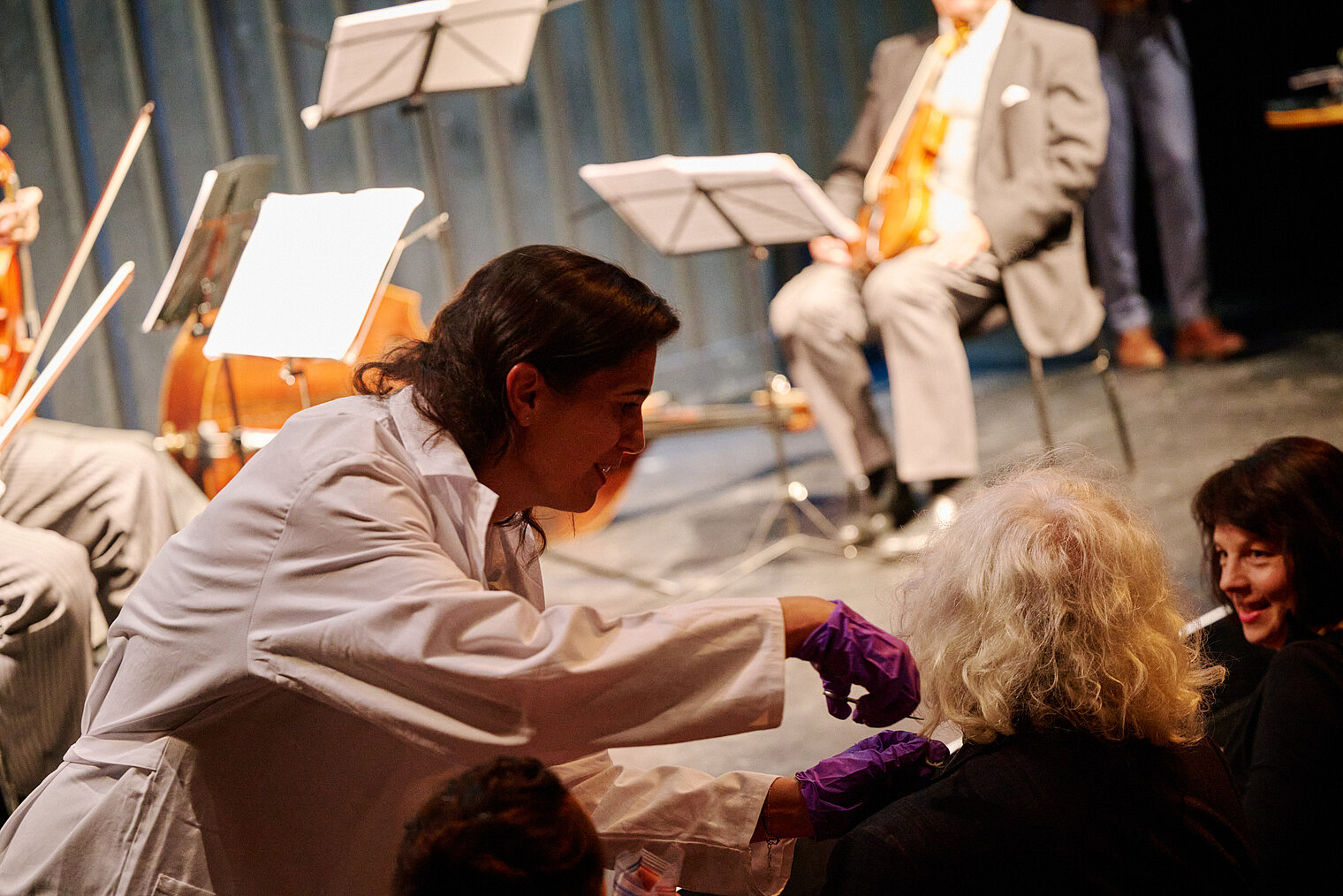
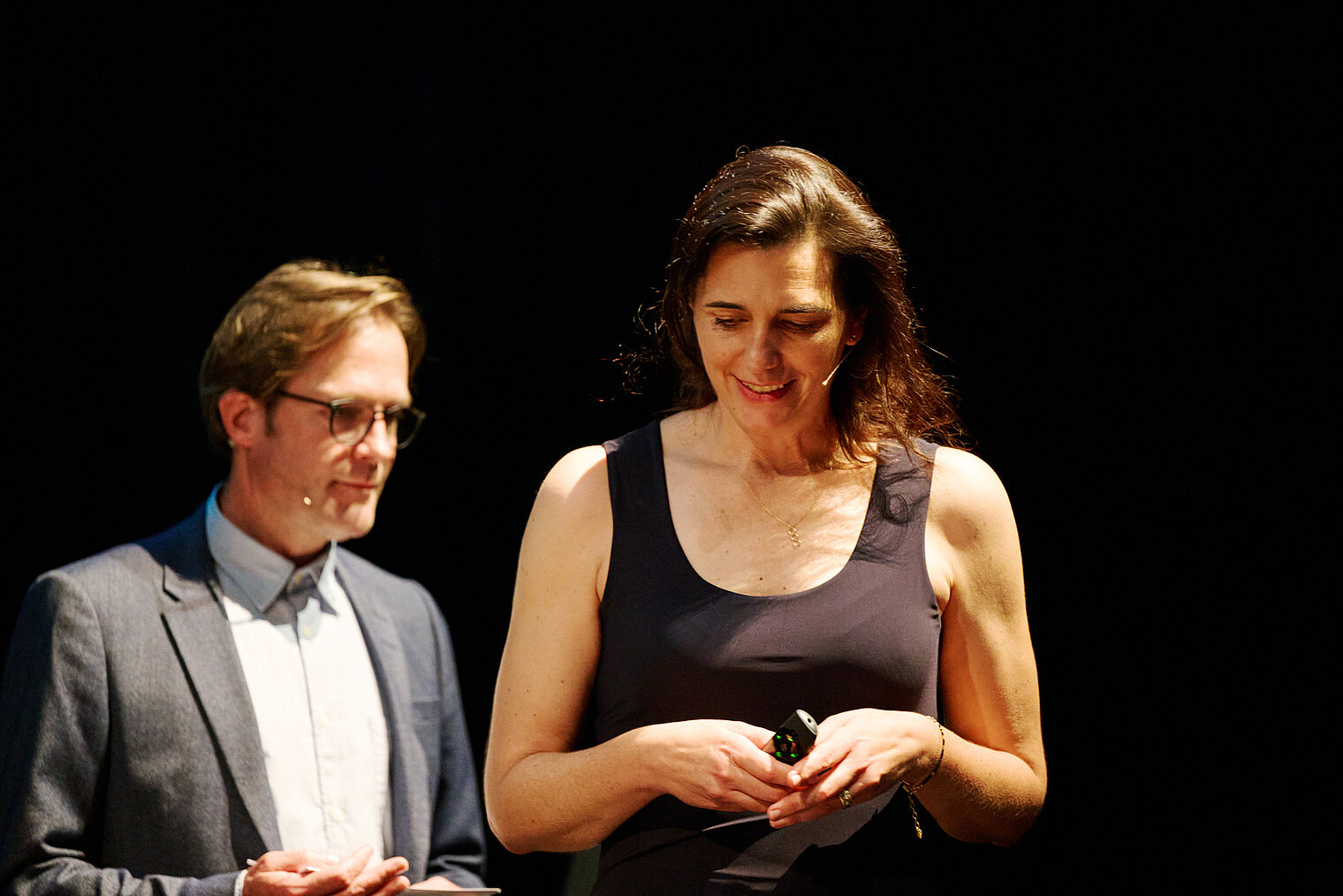
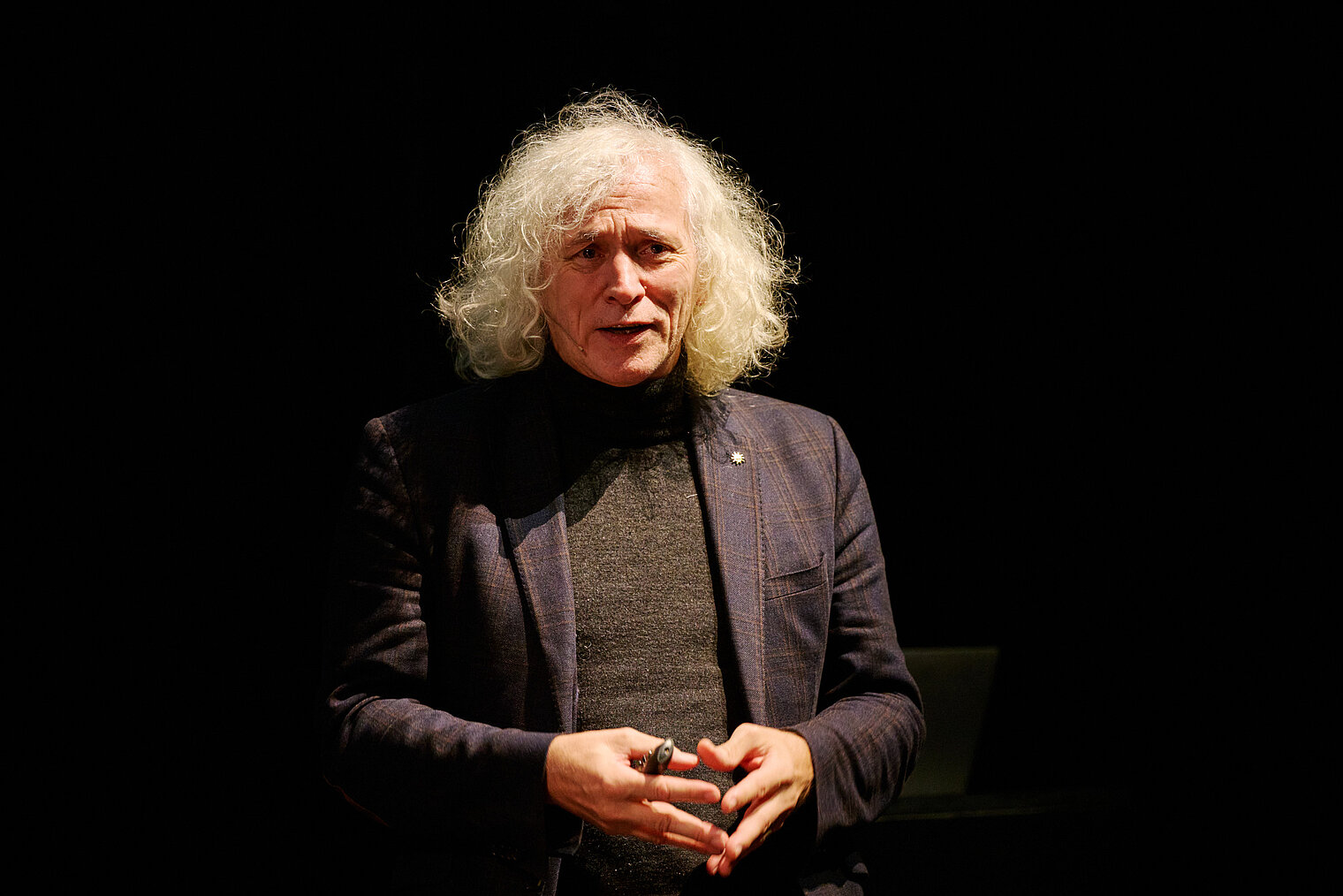
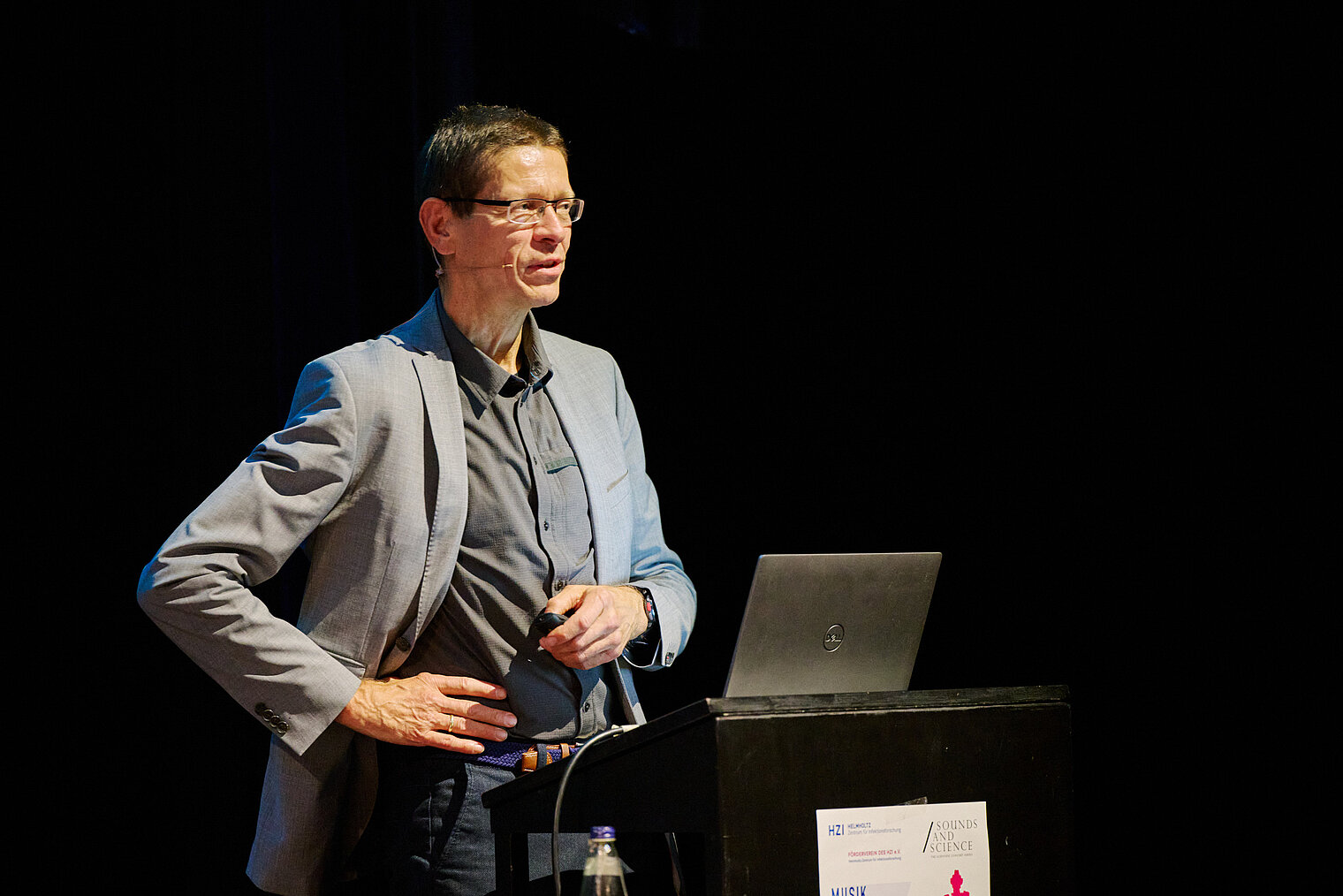
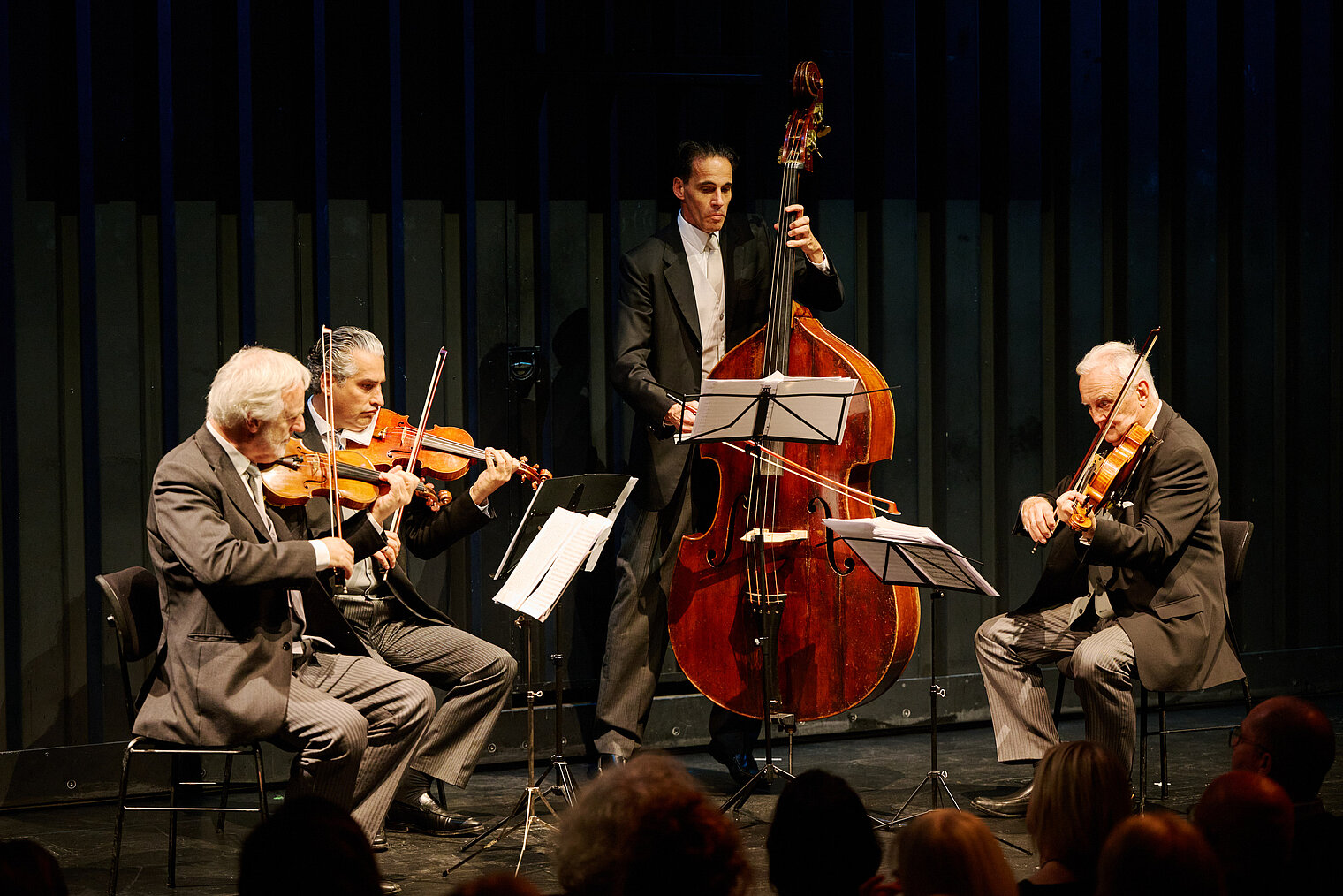
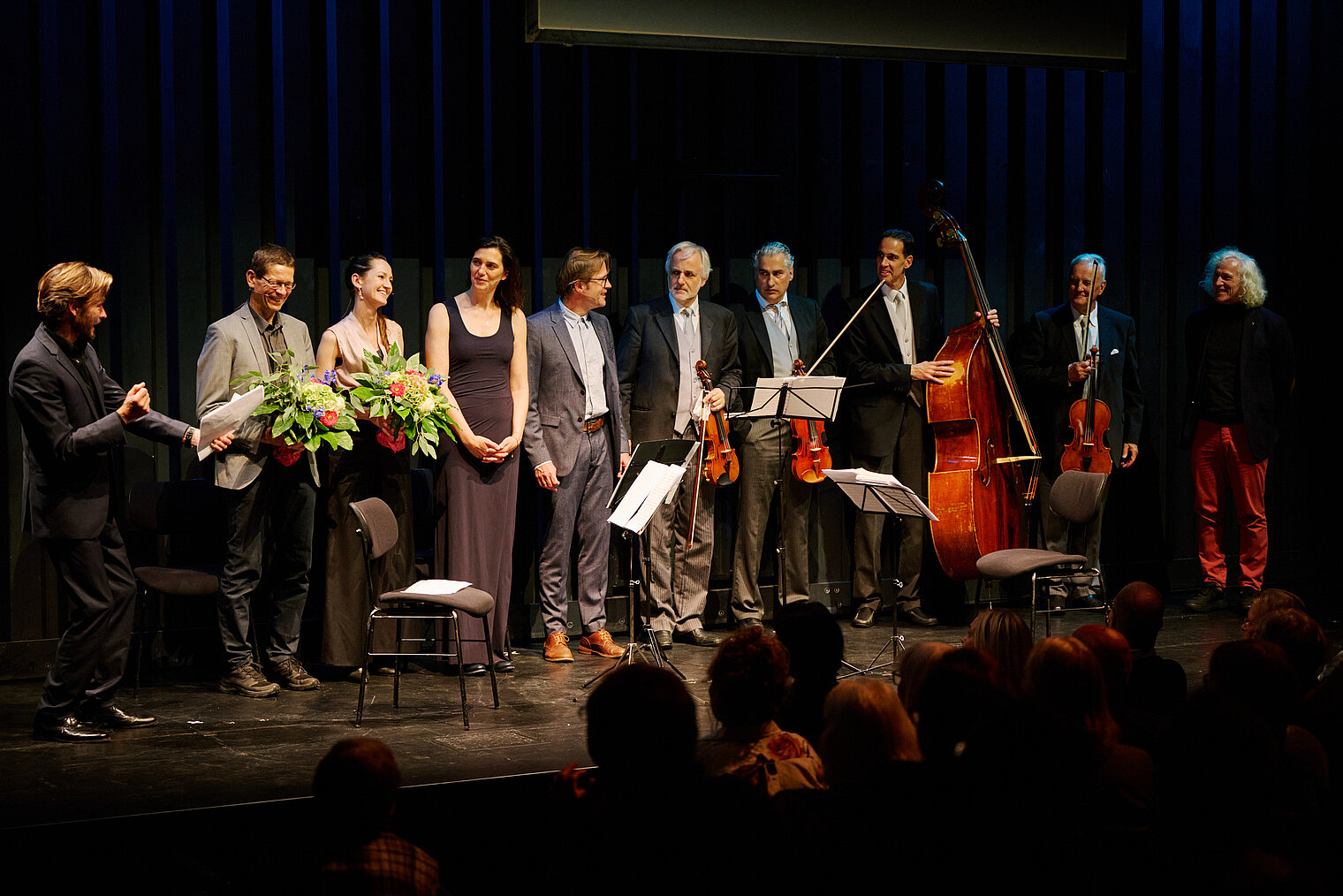
Am 25. Oktober 2024 lud das HZI zu einem besonderen Konzert im kleinen Haus des Staatstheaters Braunschweig ein. Beim Wissenschaftskonzert „Sounds and Science: Musik und Infektionen“ brachten die HZI-Forscher:innen Kathrin de la Rosa, Melanie Brinkmann, Thomas Pietschmann, Josef Penninger und Martin Korte dem Publikum die Lebens- und Krankengeschichten berühmter Komponisten näher. Mit Themen wie dem „Lymphknoten-Orchester“, dem Fund von Hepatitis B in Beethovens Locken, der Entwicklung der Gesundheitsforschung und der positiven Wirkung von Musik auf das Gehirn schlugen die Wissenschaftler:innen eine perfekte Brücke zwischen der Musik und ihrer Forschung.
Zwischen den unterhaltsamen wissenschaftlichen Kurzvorträgen spielte ein Quartett von Mitgliedern der Wiener Philharmoniker um Rainer Honeck, Rémy Ballot, Hans Peter Ochsenhofer und Manfred Hecking Werke der vorgestellten Komponisten.
Durch den Abend führte der Moderator und Schauspieler Andreas Pietschmann.
Das Publikum zeigte sich begeistert von der einzigartigen Mischung aus Wissenschaft und Musik. Die Verbindung von wissenschaftlichen Erkenntnissen und musikalischem Genuss hinterließ einen tiefen Eindruck und sorgte für stehende Ovationen.
In der Adventszeit veröffentlichen wir wöchentlich einen Vortrag von der Veranstaltung als Podcastfolge und als Video. Sie finden die Adventsfolgen bei Podigee und auf unserem Youtube-Kanal.
Forschungspreise
Inhoffen-Medaille

Zum Gedenken an den 1992 verstorbenen Chemiker Prof. Hans Herloff Inhoffen veranstalten das Helmholtz-Zentrum für Infektionsforschung (HZI) und die Technische Universität (TU) Braunschweig seit 1994 regelmäßig die Inhoffen-Vorlesung, bei der die mit 8.000,- Euro dotierte Inhoffen-Medaille vergeben wird. Die Inhoffen-Medaille ist einer der angesehensten deutschen Preise auf dem Gebiet der Naturstoffchemie.
Inhoffen lehrte von 1946 bis 1974 an der TU Braunschweig und amtierte dort von 1948 bis 1950 als Rektor. Er gründete darüber hinaus 1965 das "Institut für Molekulare Biologie, Biochemie und Biophysik", das Vorläufer-Institut der "Gesellschaft für Biotechnologische Forschung" und damit des heutigen HZI.
Die Inhoffen-Medaille wird durch den Förderverein des HZI verliehen, der auch das Preisgeld in Höhe von 8.000,- Euro finanziert.
Der Preisträger des Jahres 2024 ist Prof. Stephan A. Sieber von der TU München.

PREISTRÄGER:INNEN INHOFFEN-MEDAILLE:
- 2024: Stephan A. Sieber, TU München
- 2023: Jörn Piel, ETH Zürich, Schweiz
- 2022: Sarah Reisman, California Institute of Technology, USA
- 2020: Christian Hertweck, Leibniz-Institut für Naturstoff-Forschung und Infektionsbiologie – Hans-Knöll-Institut (HKI) Jena
- 2019: Phil Baran, The Scripps Research Institute, La Jolla, USA
- 2018: Rolf Müller, HIPS, Helmholtz-Institut für Pharmazeutische Forschung Saarland
- 2017: Helma Wennemers, ETH Zürich, Schweiz
- 2016: Thomas Carell, Ludwig-Maximilians-Universität München
- 2015: Hiroyuki Osada, RIKEN Center for Sustainable Resource Science, Japan
- 2014: Alois Fürstner, Max-Planck-Institut für Kohlenforschung
- 2013: Christopher T. Walsh, Harvard Medical School, USA
- 2012: Peter Leadlay, Abteilung für Biochemie, Universität Cambridge, GB
- 2011: Peter Seeberger, Max-Planck-Institut für Kolloid- und Grenzflächenforschung, Potsdam
- 2010: Herbert Waldmann, Max-Planck-Institut für Molekulare Physiologie, Dortmund
- 2009: William H. Fenical, Scripps Institution of Oceanography, USA
- 2008: Steven V. Ley, Universität Cambridge, GB
- 2007: François Diederich, ETH Zürich
- 2006: Gerhard Höfle und Hans Reichenbach, Helmholtz-Zentrum für Infektionsforschung, Braunschweig
- 2005: Wilhelm Boland, Max-Planck-Institut, Jena
- 2003: Manfred T. Reetz, Mülheim an der Ruhr
- 2002: Horst Kessler, Universität München
- 2001: Pierre Potier, CNRS, Gif-sur-Yvette, Frankreich
- 2000: Rudolf Wiechert, Berlin
- 1999: Carl Djerassi, Universität Stanford, USA
- 1998: Ekkehard Winterfeldt, Universität Hannover
- 1997: Sir Alan R. Battersby, Universität Cambridge, GB
- 1996: Kyriacos C. Nicolaou, Scripps Institute, La Jolla, USA
- 1995: Albert Eschenmoser, ETH Zürich
- 1994: Gerhard Quinkert, Universität Frankfurt
Jürgen Wehland-Preis

Der Jürgen Wehland-Preis wird zu Ehren des ehemaligen wissenschaftlichen Geschäftsführers des HZI verliehen, der im Jahr 2010 nach nur einjähriger Amtszeit unerwartet verstarb. 2023 wurde der Preis zum neunten Mal gemeinsam vom HZI und dem Förderverein des HZI vergeben.
Die Virologin Jun. Prof. Stephanie Pfänder hat den Jürgen Wehland-Preis 2023 erhalten. Sie ist Wissenschaftlerin an der Ruhr-Universität Bochum in der Abteilung für Molekulare und Medizinische Virologie und beschäftigt sich in ihrer Forschung mit sogenannten Emerging Viruses, insbesondere mit Coronaviren. Der Preis wurde ihr im feierlichen Rahmen und im Beisein der Braunschweiger Bürgermeisterin Anke Kaphammel sowie Gästen aus Wissenschaft und dem Alumni-Netzwerk des Helmholtz-Zentrums für Infektionsforschung (HZI) verliehen. Das HZI vergibt den mit 5000 Euro dotierten Jürgen Wehland-Preis bereits zum neunten Mal und ehrt damit Nachwuchswissenschaftler:innen, die sich auf dem Gebiet der Infektionsforschung verdient gemacht haben. Die Preisverleihung fand im Veranstaltungszentrum HZI Forum in Braunschweig statt.
PREISTRÄGER:INNEN DES JÜRGEN WEHLAND-PREIS:
- 2023: Jun. Prof. Stephanie Pfänder, Ruhr-Universität Bochum
- 2020: Dr. Kilian Schober, TU München
- 2018: Dr. Katherine Beckham, EMBL European Molecular Biology Laboratory Hamburg
- 2016: Dr. Luciana Berod, TWINCORE Hannover
- 2015: Dr. Sabrina Schreiner, TU München und Helmholtz-Zentrum München
- 2014: Prof. Andreas Müller, Otto-von-Guericke Universität Magdeburg
- 2013: Dr. Andrea Ablasser, Universität Bonn
- 2012: Dr. Stephanie Bertram, Deutsches Primatenzentrum
- 2011: Dr. Eike Steinmann, TWINCORE Hannover
Promotionspreis

Der Förderverein des HZI vergibt jährlich zwei Promotionspreise zur Förderung des wissenschaftlichen Nachwuchses. Die Auszeichnung würdigt hervorragende Dissertationen aus dem Bereich der Lebenswissenschaften, die an einer Partnerhochschule des HZI abgeschlossen wurden.
Die Promotionspreise sind jeweils mit 1.000 Euro dotiert und werden im Rahmen der jährlich stattfindenden Hans Herloff Inhoffen-Vorlesung verliehen.
Die Preisträger für Promotionen, die im Vorjahr abgeschlossen wurden, werden nach Eingang der Bewerbungen ausgewählt.
PREISTRÄGER:INNEN DER PROMOTIONSPREISE:
- 2024: Dr. Matthias Bruhn, Dr. Blondelle Matio Kemkuignou
- 2023: Dr. Alaa Alhayek, Dr. Chunlei Jiao, Dr. Bernard C. Silenou
- 2022: Dr. Chantal Bader, Dr. Birthe Reinecke
- 2021: Dr. Silke Rath, Dr. Matthias Schaks
- 2020: Dr. Adrian Kordes, Dr. Markus Stempel
- 2019: Dr. Katharina Borst, Dr. Andreas Martin Kany
- 2018: Dr. Frieda Kage, Dr. Daniel Todt
- 2017: Dr. Tobias Bock, Dr. Islam Ahmed Mohamed El-Awaad
- 2016: Dr. Stephanie Pfänder, Dr. Eric Kuhnert
- 2015: Dr. Michaela Annemann, Dr. Michael Storz
- 2014: Dr. Stefanie Kristin Wöhl-Bruhn, Dr. Christian A. Citron
- 2013: Dr. Cornelia Chizzali, Dr. Christian Mayer
- 2012: Dr. Christina Ziegler, Dr. Nick Quade
- 2011: Dr. Julia Bitzegeio, Dr. Leonor da Gama Cavalho Norton
- 2010: Dr. Judith Becker, Dr. Jan Hänisch
- 2009: Dr. Jacek Puchalka
- 2008: Dr. Thomas Wollert, Dr. Pablo Becker
- 2007: Dr. Roland Adden, Dr. Raimo Franke
- 2006: Dr. Silke Wenzel, Dr. Annika Steffen
- 2005: Dr. Oliver Goldmann, Dr. Jibin Sun
- 2003: Dr. Simone Bergmann, Dr. Philipp Hartmann
- 2002: Dr. Nicole Glaser, Dr. Edelweiß Markworth, Dr. Andreas Toman
- 2001: Dr. Oliver Pabst, Dr. Stefan Hüttelmaier
- 2000: Dr. Armin Bauer, Dr. Ursula Deiters
Paper of the Month
Ein internes Gremium wählt monatlich für den HZI Paper of the Month Award (POM) eine Publikation aus. Der Preis ist ein Geschenkgutschein für den Erstautor/die Erstautorin, begleitet von einer Urkunde. Die Verleihung findet regelmäßig im Anschluss an das HZI-interne Friday Lunch Seminar statt.
Aktuelle Preisträger:innen
Januar 2024
Chengzhang Fu (HIPS), Yunkun Liu (HIPS), Christine Walt (HIPS), Sari Rasheed (HIPS)
Elucidation of unusual biosynthesis and DnaN-targeting mode of action of potent anti-tuberculosis antibiotics Mycoplanecins
doi: 10.1038/s41467-024-44953-5
Februar 2024
Svenja M. Sake (Twincore)
Drug repurposing screen identifies lonafarnib as respiratory syncytial virus fusion protein inhibitor
doi: 10.1038/s41467-024-45241-y
März 2024
Milan Gerovac (HIRI)
Phage proteins target and co-opt host ribosomes immediately upon infection
doi: 10.1038/s41564-024-01616-x
April 2024
Javier Botey (CIIM, Twincore)
A comprehensive genetic map of cytokine responses in Lyme borreliosis
doi: 10.1038/s41467-024-47505-z
Mai 2024
Lisa Osbelt-Block (HZI)
Klebsiella oxytoca inhibits Salmonella infection through multiple microbiota-context-dependent mechanisms
doi: 10.1038/s41564-024-01710-0
Juni 2024
Ronald Garcia (HIPS), Alexander Popoff (HIPS), Chantal Bader (HIPS)
Discovery of the Pendulisporaceae: An extremotolerant myxobacterial family with distinct sporulation behavior and prolific specialized metabolism
doi: 10.1016/j.chempr.2024.04.019